Service Line:+86-022-82164980
Address:FL-4, Building A5, International Enterprise Community, Tianjin, China
Email:[email protected]
The primary sequence of protein determines the advanced structure of a protein, influencing a protein's function. Therefore, the sequence of a protein is essential for understanding the biological function of the protein. KMD Bioscience has developed a comprehensive protein sequencing platform to meet the diverse needs of our customers, including N-terminal sequencing and de novo protein sequencing. Welcome to contact us for more information.
Protein Sequencing Service
Protein sequence is an important physical and chemical property of proteins, which is of great significance for the identification of protein species and advanced structure prediction. With the development and progress of life science technology, many protein sequencing technologies have emerged, which can be roughly divided into mass spectrometry and non-mass spectrometry. Non-mass spectrometry mainly includes the Sanger method, Edman degradation method, dinitrofluorobenzene method, dan sulfonyl chloride method, hydrazine hydrolysis method, and carboxypeptidase method, among which the Edman degradation method is the most commonly used non-mass spectrometry method.
Both mass spectrometry and Edman degradation can be used for protein sequence analysis, but there are great differences in the analysis principle and the sequence region that can be measured. Edman degradation method, a chemical method, which is a sequencing method for N-terminal amino acids. It does not need to rely on a protein database to directly analyze the data and directly obtain the amino acid sequence. Through three successive chemical reactions, an N-terminal amino acid residue can be cut off, and the amino acid is identified, and then the protein with one amino acid missing is carried out the same operation again, and so on, identifying one amino acid residue each cycle. However, end-modified or blocked amino acids cannot carry out the above chemical reactions, so the Edman degradation method is powerless to N-terminal modified or blocked proteins. The quality of Edman sequencing decreases as the number of amino acids detected increases, so it is usually only used to identify the first 30-60 amino acids, and its detection length does not exceed 100 amino acids. Therefore, Edman sequencing cannot be used for full-length protein sequencing.
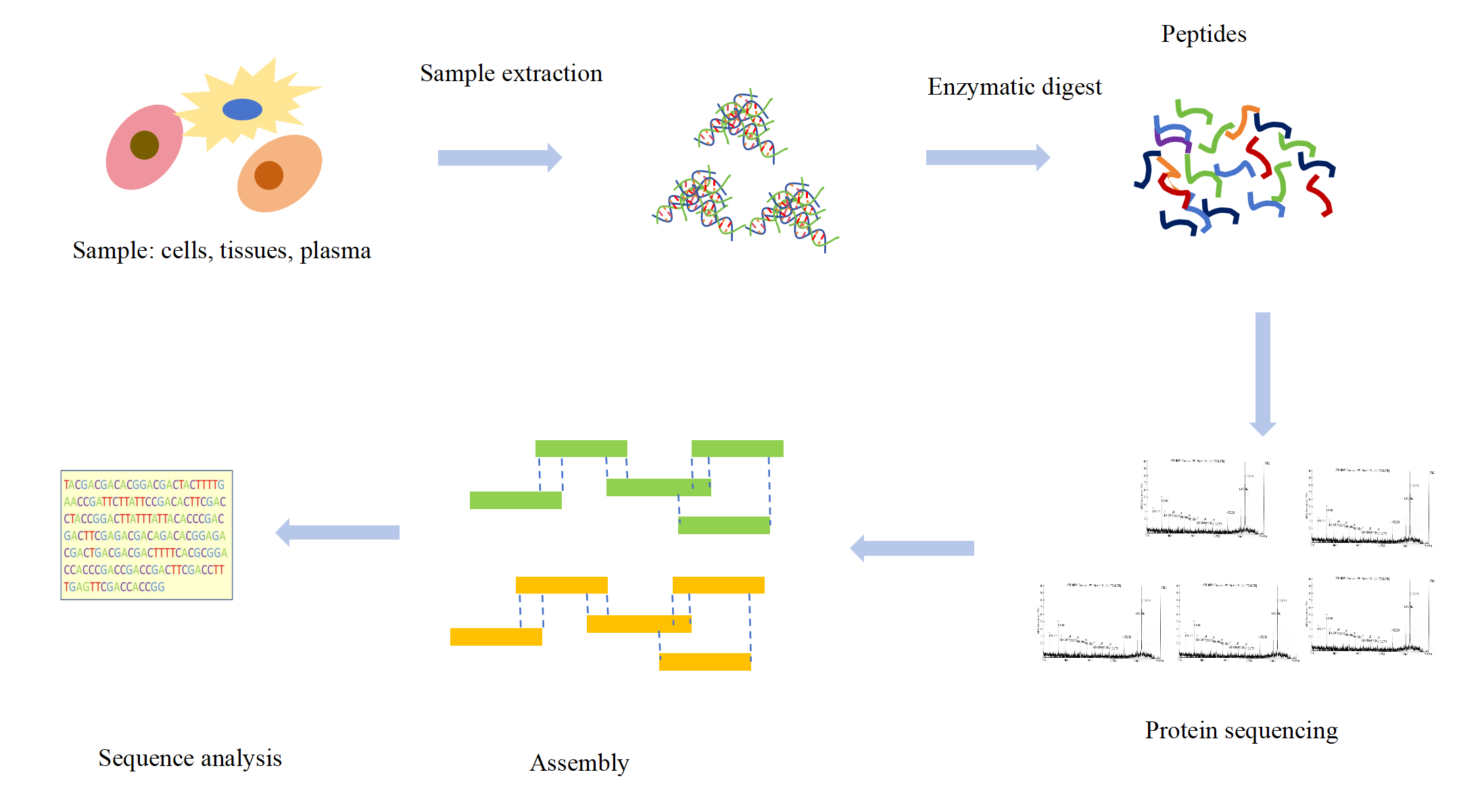
The protein is digested into a peptide of 5-25 amino acids by mass spectrometry, then analyzed and detected by a mass spectrometer, and the collected data is matched with the theoretical sequence database to confirm the N-terminal sequence. Mass spectrometry, which can detect the closed and modified N-terminus of proteins, has become the preferred method for full-length antibody sequencing. Compared with the Edman degradation method, mass spectrometry can realize the full-length sequence analysis of proteins, and can also be used for the sequence analysis of N-terminal modified proteins, breaking the limitations of the Edman degradation method. In addition, mass spectrometry can also perform de novo sequencing, providing a solution for the determination of a completely new or no theoretical database of protein sequences.
KMD Bioscience has a high-quality Orbitrap Fusion Lumos mass spectrometer, established a perfect database combination sequencing service, and can provide our customers with high-quality protein sequencing services. Given different experimental needs, our company also provides the Edman degradation method for the determination of protein N-terminal up to 67 amino acid sequence length.
De Novo Protein Sequencing Service
Based on the advanced mass spectrometry instrument, 'Dao Jin' PPSQ-31A, and combined with rich experience in bioinformatics analysis, KMD Bioscience has established a completely new generation of de novo protein sequencing platform that can achieve accurate sequencing analysis of monoclonal antibodies and proteins. After the sample is received our scientists will first identify and fragment it with six commonly used proteases: Trypsin, Chymotrypsin, Asp-N, Glu-C, Lys-C, and Lys-C. KMD Bioscience is working to get the most fragmented sequences to achieve 99.99% protein coverage. The sequence information was spliced by PEAKs Studio and PEAKs Ab software and then artificially joined to perform nearly 99.99% correct protein or antibody primary sequence.
For more de novo protein sequencing services, please browse the antibody de novo sequencing services or contact us for further information.

Fig.1 Orbitrap Fusion Lumos
Orbitrap Fusion Lumos Mass spectrometer is a three-in-one combined mass spectrometer, which is equipped with a new atmospheric pressure ion source, advanced quadrupole technology, ultra-high-field Orbitrap analyzer, and the latest two-pressure linear ion trap, which is the most advanced mass spectrometer. Orbitrap Fusion Lumos is designed to expand proteomics, biopharmaceutical, and metabolomics applications through advanced performance, including applied isotope labeling quantification, low-level P-analysis, data-independent acquisition (DIA), and top-down proteomics studies. Due to its excellent performance and advanced configuration, it can be applied to many aspects, such as high-throughput protein identification, large-scale proteomic non-label quantitative analysis, protein post-translational modification PTMs analysis (such as acetylation, ubiquitination, phosphorylation, glycosylation, etc.) and protein top-down analysis.
N-Terminal Sequencing Service (Edman Degradation)
KMD Bioscience can sequence up to 60 amino acid residues from the N-terminal. The Edman method for sequencing a single amino acid cycle consists of 3 steps.
Step 1: PITC binds to free amino acids at the N-terminal of the protein under alkaline conditions.
Step 2: N-terminal residues are removed in acidic solutions.
Step 3: PITC-bound residues are converted to more stable PTH residues. After online analysis by HPLC, amino acids were determined according to elution time.
Protein Sequencing Process
Protein samples are first separated by SDS-PAGE to ensure the purity can meet the requirements of sequencing. Subsequently, protein samples from SDS-PAGE are transferred to PVDF membranes, and cut the gel band after dyeing. Then it is loaded directly on the sequencer.
Sample Requirement For Protein Sequencing
|
Services Available |
Sample Type |
Sample State |
Standard Sample Size |
Purity Requirement |
Attention |
|
De novo protein sequencing |
monoclonal antibody; Antibody-drug conjugate; recombinant protein; Polypeptide drug |
Liquid/dry powder/protein adhesive strip |
≥100ug |
≥90% |
1. Try not to contain BSA in the sample. If there is BSA, make a column in the buffer to note the detailed concentration. 2. SDS-PAGE results of recombinant protein samples should be provided to provide information on whether they are dimerized or polymerized. 3. N-terminal sequencing, C-terminal protein sequencing, and protein sequence analysis need to provide reference sequences. 4. The sample size of each hole should be at least 15 micrograms, each sample should be sent to at least 6 glue holes, stored at 4℃, placed in an EP tube, soaked in deionized water, and sent in an ice pack or dry ice. 5. The protein solution buffer can be pure water or PBS buffer. If there are other solution components, please indicate them in the sample information. The prepared samples were stored at -20℃, placed in EP tubes, and sent in ice packs or dry ice. 6. The protein powder is freeze-dried from pure water or PBS buffer, and contains other solution ingredients, please indicate in the sample information. The prepared samples were put into EP tubes and sent at room temperature or in ice packs; 8. Prior communication is required before sending samples. |
|
Peptide coverage analysis |
≥100ug |
≥90% |
|||
|
N-terminal protein sequence analysis |
≥100ug |
≥90% |
Protein Sequencing Service Characteristics
-- Identify recombinant protein expression and pulldown products.
-- Validation of expressed products in cell lines.
-- Validation of N-terminal sequences of biosimilar antibody.
Protein Sequencing Service Highlights
--Unparalleled protein sequencing accuracy.
--99.99% protein sequence coverage and confidence.
--Strict quality control system and strong analytical capabilities.
How to Order?
If you have any questions regarding our services or products, please feel free to contact us by E-mail: [email protected] or Tel: +86-400-621-6806